|
 |
 |

|

|
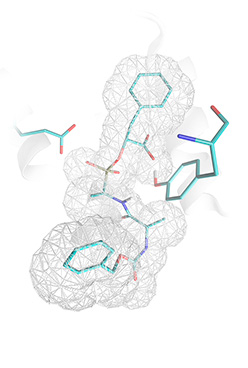
CarboxypeptidaseA inhibitor successfully docked by Lead-Finder. A — crystallographic pose (PDB structure 6cpa), B — ligand pose predicted by Lead-Finder. |
Docking success rateDocking success rate was benchmarked as a percentage of correctly docked ligands (for which top-scored pose was within 2 Å RMSD from the reference ligand coordinates) for a set of protein-ligand complexes extracted from PDB. Though alternative definitions of docking success rate are available1 we use the most common one, to make Lead-Finder benchmarks comparable with competitive software results obtained elsewhere. Additionally, for the sake of reproducibility all docking calculations are performed independently 20 times, and only if probability of generating top-ranked pose with 2 Å accuracy is greater or equals 0.5; docking is recognized successful. A set of 407 protein-ligand complexes was used for current docking success rate measurements. This set of complexes was combined from test sets used in original benchmarking studies of such docking programs as: FlexX2, Glide SP 3, Glide XP 4, Gold 5,6,7, LigandFit 8, MolDock 9, Surflex 10. For this reason, current benchmarking study of Lead-Finder docking success rate is the most extensive study of such kind; moreover, results obtained with Lead-Finder can be fairly compared to competitive docking programs of interest, since they were obtained on the same sets of protein-ligand complexes. All benchmarking calculations were performed in two regimes (default docking and screening, see Technology section) to give impression of speed/accuracy balance. As can be seen from the table, docking success rate obtained with Lead-Finder on different test sets ranges from 80.0% (for GlideXP and FlexX test sets) to 96.0% (for Surflex and MolDock test sets). It can be seen from the table that Lead-Finder outperforms all docking programs, for which robust original benchmarks exist. More precisely, Lead-Finder successfully docks 73 structures which could not be docked by FlexX, 52 structures — for Glide SP, 52 structures — for Glide XP, 30 structures — for Gold on original Gold test set, 13 structures — for Gold on Astex diversity test set, 9 structures — for MolDock, 21 structures — for Surflex. Example of carboxypeptidase A inhibitor, which could not be correctly docked by such programs as Glide 3 or FlexX 2, but was successfully docked by Lead-Finder, is illustrated by the figure.  |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Docking success rate (%) of different programs obtained on their native test sets and current Lead-Finder benchmarks in default docking and screening regimes. You can download complete description of current docking success rate benchmarking data including composition of the entire test set of 407 protein-ligand complexes by original test sets of FlexX, Glide SP, Glide XP, Gold, LigandFit, MolDock, Surflex docking programs.
Docking performance of Lead-Finder
|
a) Data for the high-resolution subset (92 structures with resolution better than 2 Å) are not provided in original publication, only overall performance (over entire CCDC/Astex test of 224 structures) is given. Docking success rate achieved by Lead-Finder in the screening regime (with ’light´ docking algorithm settings, making it faster) was also better than original results obtained with competitive programs (except for MolDock). Such results were achieved by adjusting settings of the docking algorithm to gain reasonable decrease in docking success rate (approximately 5% on average) with maximum increase in speed of calculations. Attractiveness of the screening regime for high-performance calculations is illustrated in the Speed of Calculations section. 
|
 |
 |